Fig 1. Image of Azomycin. source
Scientists found the biosynthetic pathway to the Nitroimidazole Antibiotic Azomycin. A steppingstone towards the revolution of anaerobic bacterial infection treatments.
Scientists from the University of British Columbia found the biosynthetic pathway to the Nitroimidazole antibiotic Azomycin. The enzymatic mechanism from L-Arginine to Nitroimidazole has now been proved and present to the public. Their formal paper was published online on July 17th, 2019. Nitroimidazole is an essential component for the modern antibiotics, it is crucial to know its synthetic pathway for further pharmaceutical studies. The result of their study set the stage for further development of important anaerobic antibiotic Azomycin.
What is it? Why do we need to know about this?
Nitroimidazole is an essential antibiotic specifically to treat anaerobic bacterial infections. They are widely used to treat diseases such as Amoebiasis, Parasitic infections, skin infections, diarrhea and so on. The low redox potential of anaerobic bacteria cells allowed nitroimidazole to act as the electron sink and form the radical species. The resultant radical species would induce the bacteria cells’ death by damaging their DNA. Antibiotic is the most powerful “weapon” to fight against bacterial infections. However, according to the World Health Organization, there are more than 700000 people die every year due to antibiotic resistance. Despite the several decade’s usages of Nitroimidazole antibiotics, the drug resistance of it still remains low relatively. Thus, Nitroimidazole antibiotics are increasingly used to treat multi-drug resistant bacteria as well.
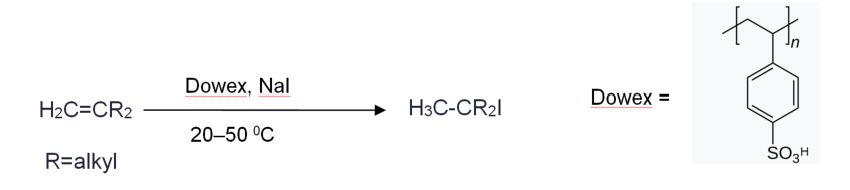
Previous research established that L-Arginine is converted to azomycin by 2-aminioimidazole. They determined that the intermediate of the reaction is 4-hydroxy-2-ketoarginine (2). Furthermore, they also observed the accumulation of pyruvate(3) side products and 2-aminoimidazole(5) from the intermediate(2). However, the actual enzymatic synthetic pathway has not been determined detailly yet. Jason and Katherine in the research group determined that PLP-dependent enzymes, RohP,RohR,RohQ and RohS plays esstential role in the catalytic pathway of the reaction. Researchers examined the in vitro activity of RohP, RohR, RohQ and RohS. They put in these enzymes separately and stepwise to different reactants. For example, in order to test whether RohR could catalyze a retro-aldol cleavage of 2 into 3 and guanidinoacetaldehyde (4), they added purified RohR instead of RohP. Then according to activity analysis and also the mass spectrum, the result shows that RohP yields a bigger portion of 2.
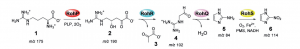
Fig 2. Reaction scheme from L-Arginine to Nitroimidazole. Source
Antibiotics are the most powerful “weapon” to kill bacteria in modern pharmaceutical studies. As early as in the 20th century, the observation of penicillin saved millions of people injured in the World War. Yet, the enormous benefits of antibiotics cause the consequences of drug-resistance. Azomycin, consider as a low-resistance antibiotic, it is crucial to understand its enzymatic reaction mechanism. Reaction mechanism allows scientist to have a more detail interpretation of the synthesis. It is crucial to find the catalytic cycle of the reaction, in order to allow scientists to develop and derive further study.
The study done by Jason and Katherine at the University of British Columbia provides the public with a steppingstone in future nitroimidazole anti-biotics study. Their study expanded people’s knowledge of the biosynthetic pathway to nitro-compounds. It also makes bacteria engineering to produce nitroaromatic compounds possible. This study will open the new door in enzymatic synthesis and biochemistry synthesis.
Cited article:
Hedges, J. B.; Ryan, K. S. In Vitro Reconstitution of the Biosynthetic Pathway to the Nitroimidazole Antibiotic Azomycin. Angewandte Chemie International Edition 2019, 58 (34), 11647–11651.